Citation
1. Zhou Z, Zou Y, Liu G, et al. Mutation-profile-based methods for understanding selection forces in cancer somatic mutations: A comparative analysis. Oncotarget 2017; 8:58835–58846.
2. Zhao W, Yang J, Wu J, et al. CanDriS: posterior profiling of cancer-driving sites based on two-component evolutionary model. Brief. Bioinform. 2021.
About
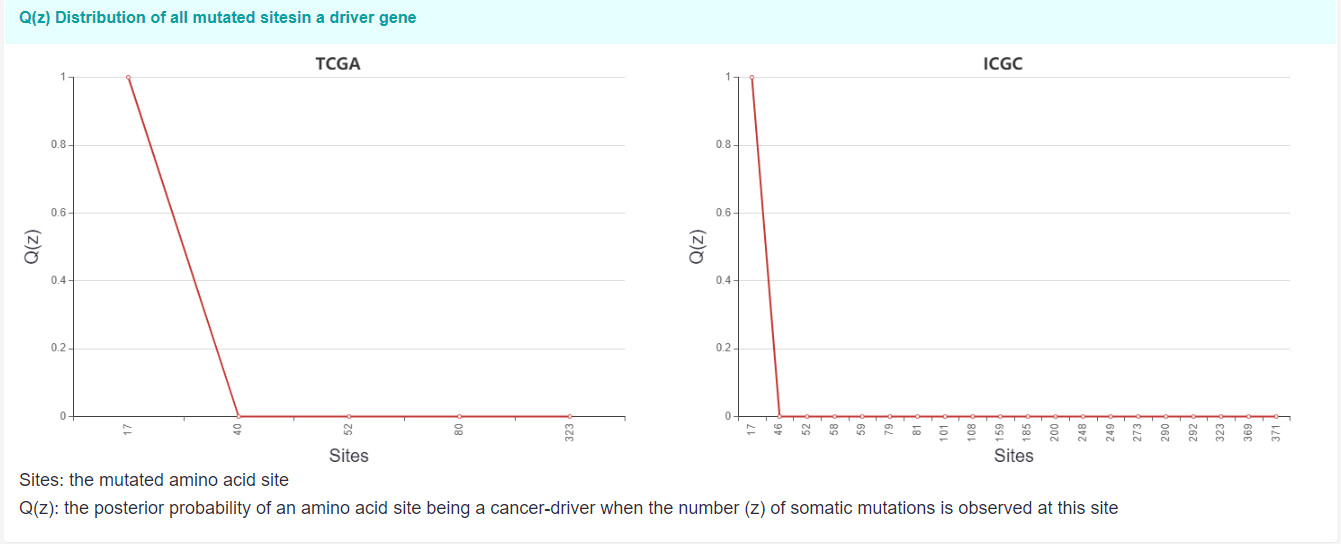
The line plot below shows the Qz distribution of all mutated sites in AKT1 in both TCGA and ICGC. (Only display the amino acid sites with at least a mutation (z ≥1)).
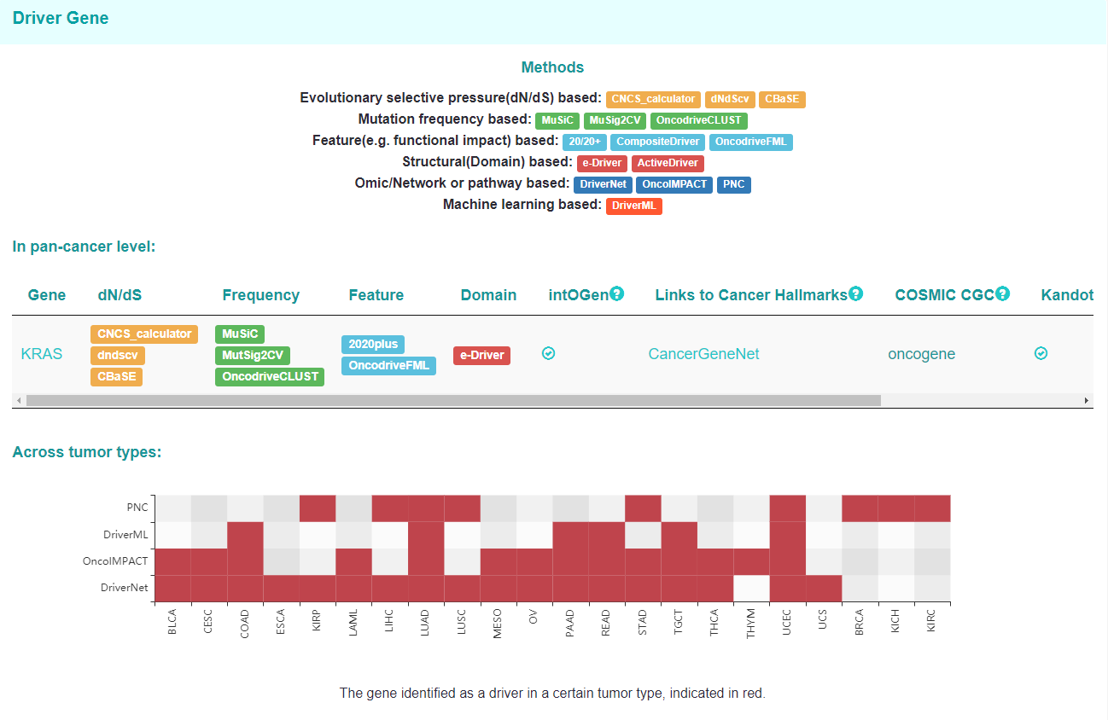

Driver Gene page
Take Gene AKT1 for example:

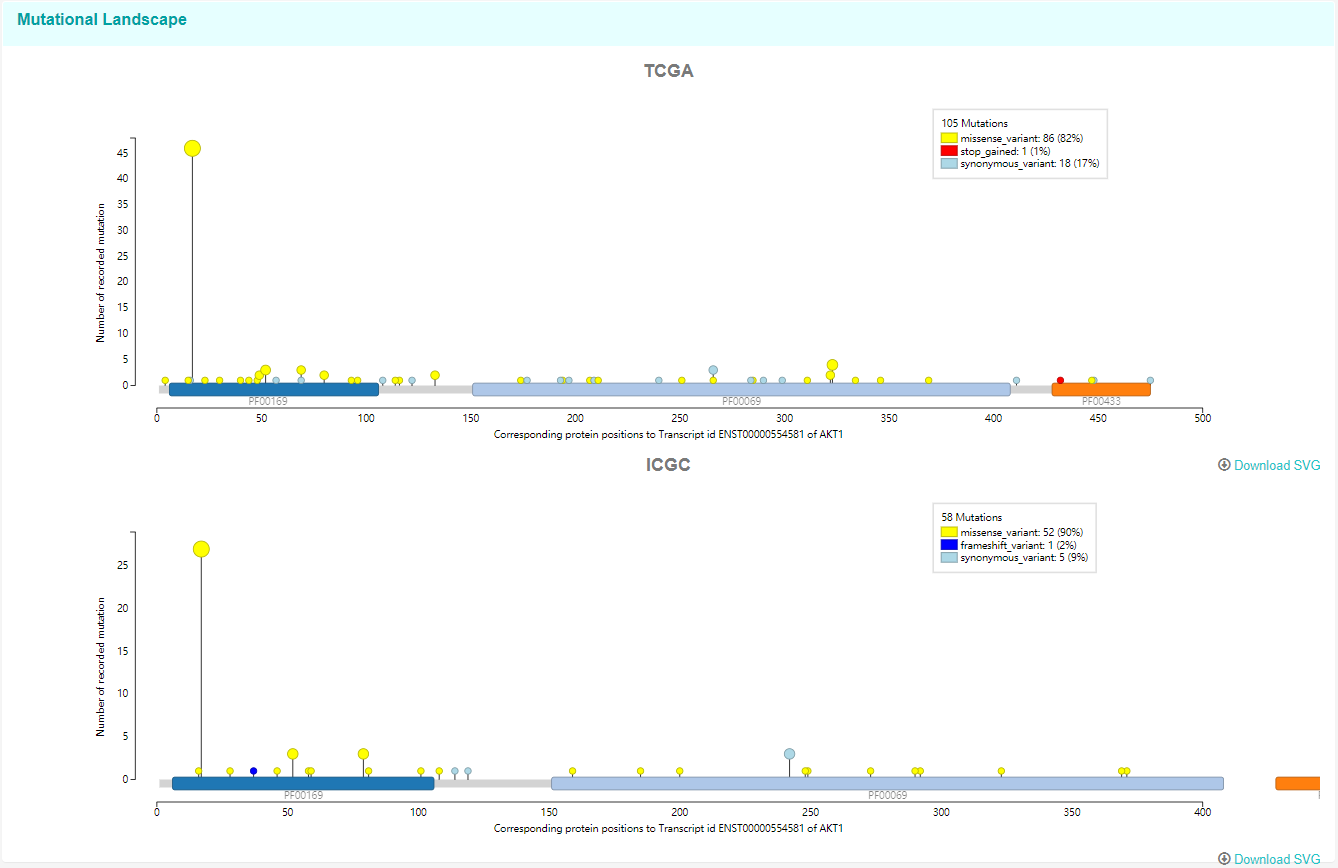
The needle plot below shows the mutational lanscape of AKT1 in both TCGA and ICGC.

The line plot below shows the Qz distribution of all mutated sites in AKT1 in both TCGA and ICGC. (Only display the amino acid sites with at least a mutation (z ≥1)).
Sites: the mutated amino acid site.
Q(z): the posterior probability of an amino acid site being a cancer-driving when the number (z) of somatic mutations is observed at this site.

These part shows different types of methods as well as some curated lists of driver genes which identify AKT1 as a driver gene in pan-cancer level and across different tumor types.

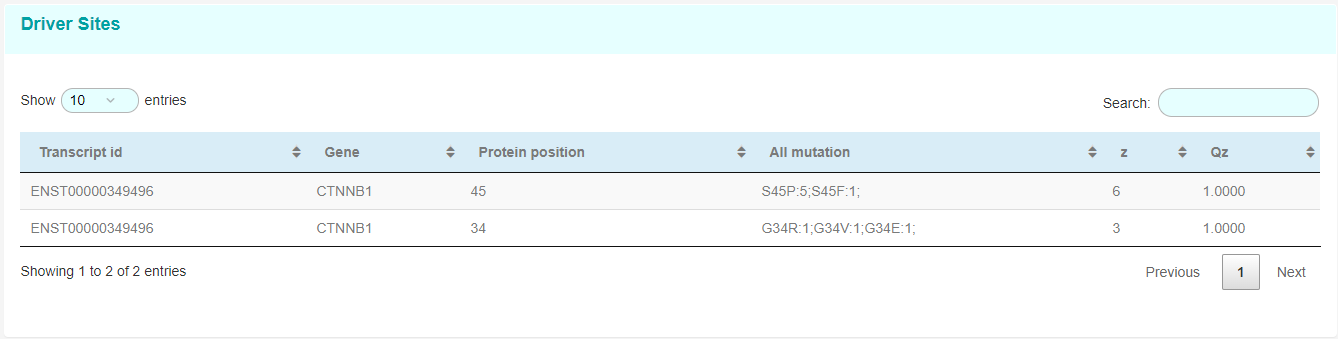
Driver Site page
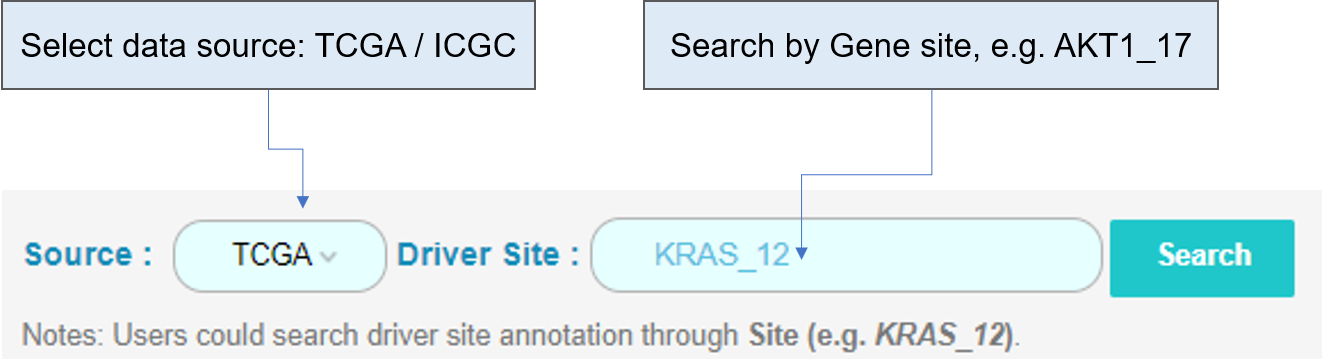
Take Site AKT1_17 for example:

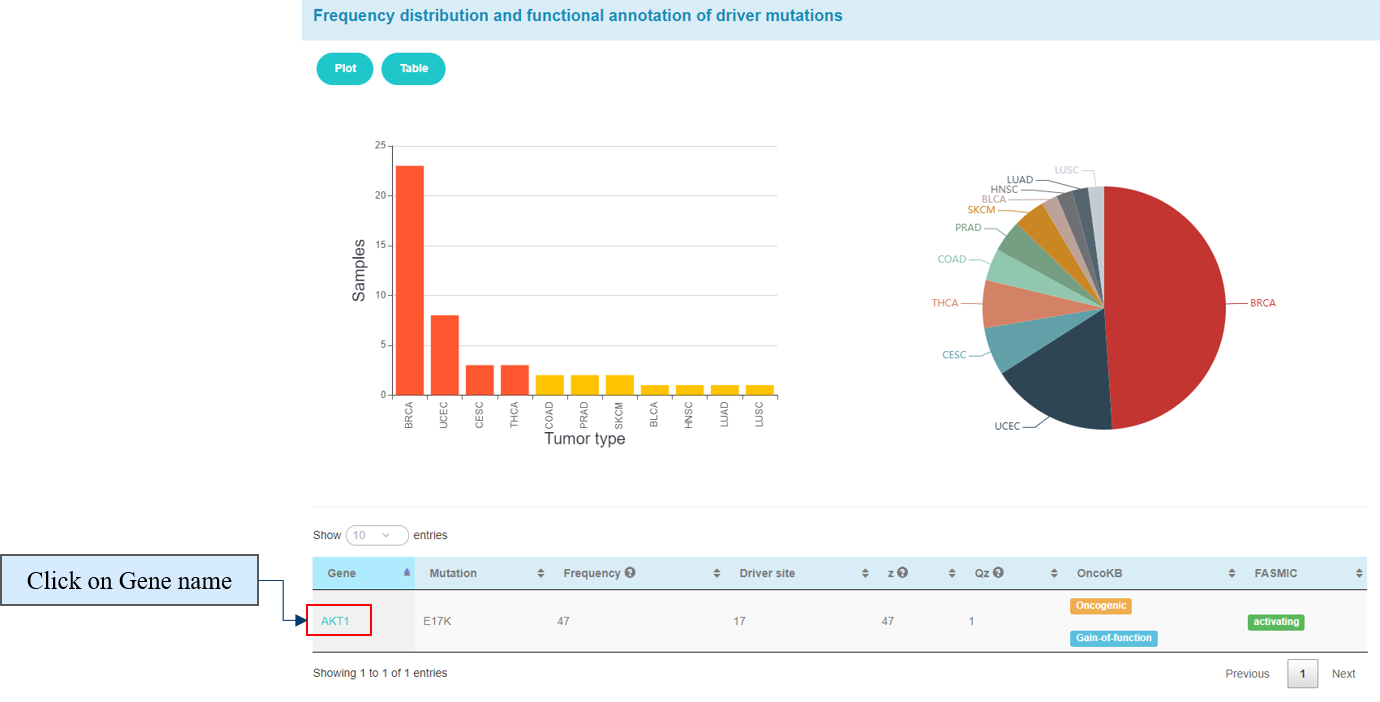
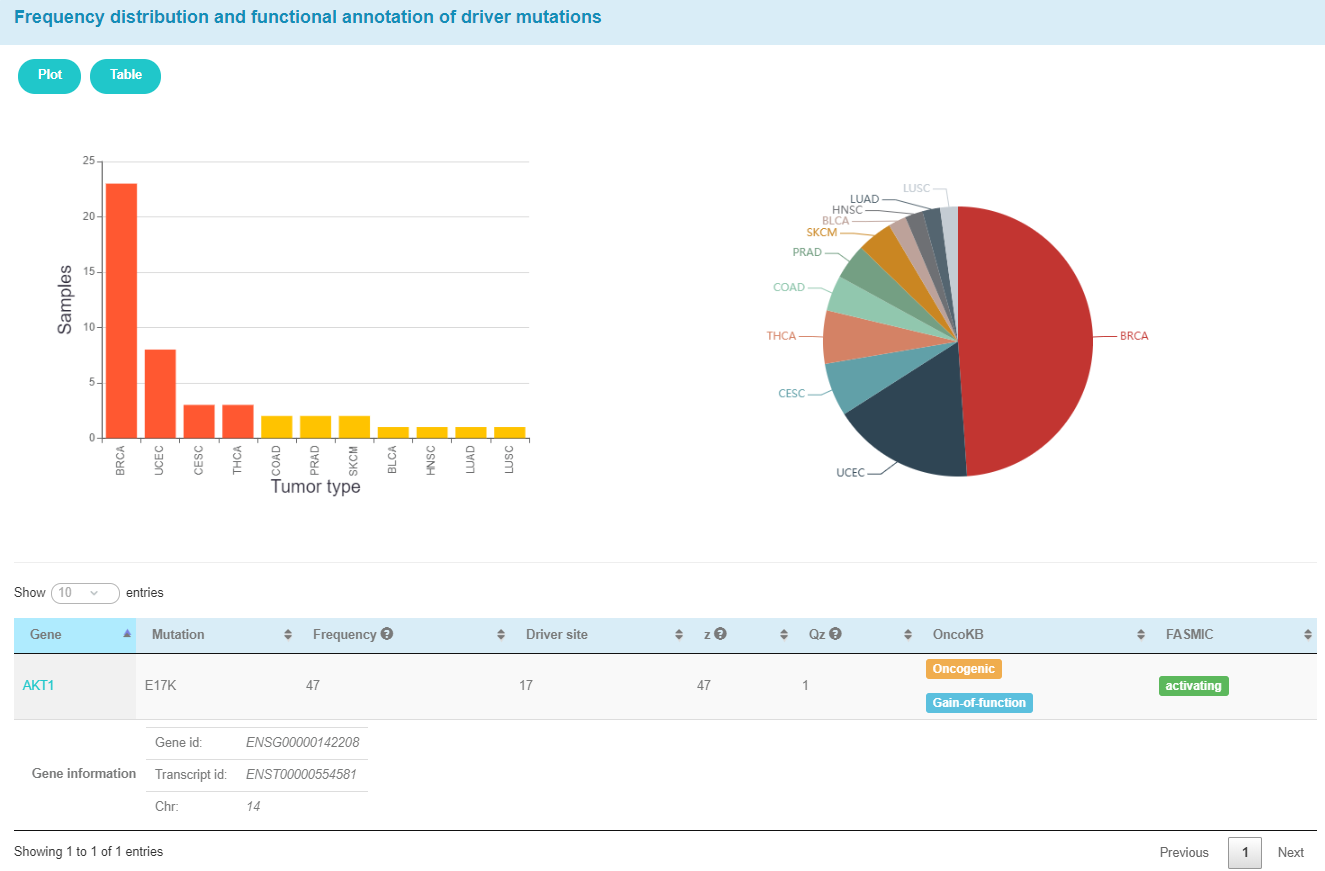
The pie plot and bar plot below show the frequency of AKT1_17 across different tumor types. The table shows functional annotation of AKT1_17 and some parameters calculated by our in-house method CanDriS.

The child table shows some information about AKT1.

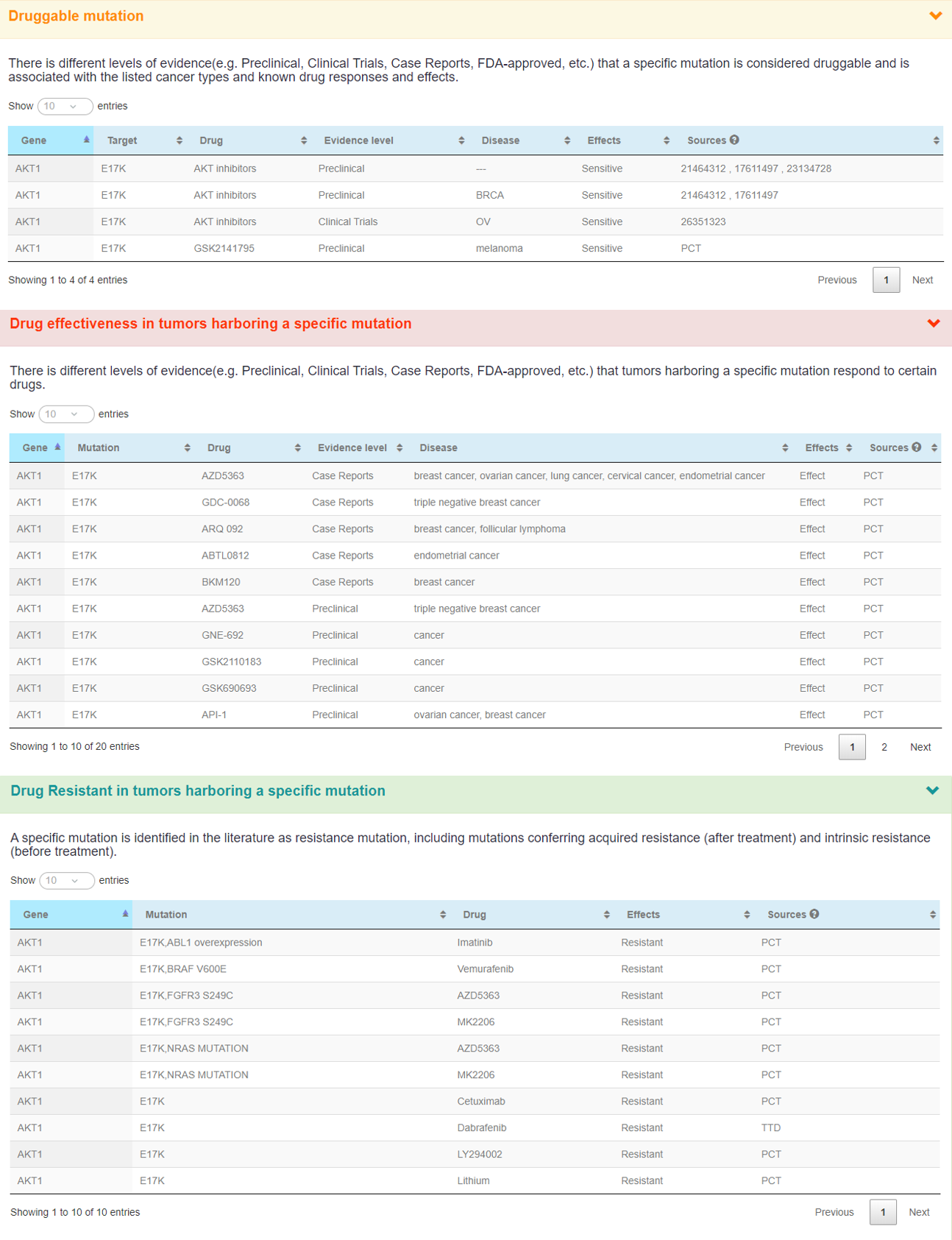
The following panels display some pharmacogenomic annotation information about AKT1_17.

Tumor Type page
Take Tumor type Adrenocortical carcinoma (ACC[TCGA]) for example:

The following table displays driver sites in Adrenocortical carcinoma (ACC[TCGA]).

Data
Tumor mutation data
Tumor somatic mutations data is collected from TCGA PanCanAtlas project and ICGC Release 25.
Driver site annotation dataMost of functional annotation data is collected from OncoKB and FASMIC.
Most of pharmacogenomics annotation data is collected from DrugBank, PharmGKB, DEPO, PCT, COSMIC Drug Resistance and TTD.
Tools
Driver gene identification methods
CNCS calculator: an in-house method to profiling Cancer driver genes by calculating Cancer-specific selection pressure.
dNdScv: a suite of maximum-likelihood dN/dS methods designed to quantify selection in cancer and somatic evolution.
CBaSE: a tool which derives gene-specific probabilistic estimates of the strength of negative and positive selection in cancer.
MuSiC: a frequency based tool used to identify significantly mutated genes.
MutSig2CV: a method analyzes somatic point mutations discovered in DNA sequencing, identifying genes mutated more often than expected by chance given inferred background mutation processes.
20/20+: a Random Forest machine learning algorithm for predicting oncogenes and tumor suppressor genes from somatic mutations.
CompositeDriver: a novel computational method considering both mutation recurrence and functional impact of mutations to identify signals of positive selection.
OncodriveFML: a method dentifies genes that have greater accumulation of mutations that have higher predicted function impact (functional impact bias).
OncodriveCLUST: a method identifies genes with non-silent mutations that cluster together in protein sequence more than expected based on a background distribution of synonymous mutations.
e-Driver: a method identifies protein regions that are enriched in somatic missense mutations using a binomial test and assuming mutations are distributed randomly across the protein.
ActiveDriver: a method detects genes that are enriched in somatic mutations located in post-translationally modified sites, such as phosphorylation, acetylation, or ubiquitination sites.
DriverNet: a package to predict functional important driver genes in cancer by integrating genome data (non-synonymous SNVs, indels, and copy number alteration) and transcriptome data (gene expression data).
OncoIMPACT: a model-driven approach to integrate omics profiles (genomics and transcriptomics) and provides patient-specific cancer driver gene predictions.
PNC: a novel network control model for identifying personalized driver genes in cancer.
DriverML: a machine learning algorithm for identifying driver genes in cancer sequencing studies.
Driver site identification method
CanDriS: an in-house method to profiling cancer-driving sites by calculating posterior probability.
Visualization
Echarts has been used for various plots and Mutations Needle Plot library has been for the mutation distribution plots.
Contact
E-mail: zhanzhou@zju.edu.cn.
E-Address: College of Pharmaceutical Sciences, Zhejiang University, Hangzhou, China.
Term of use